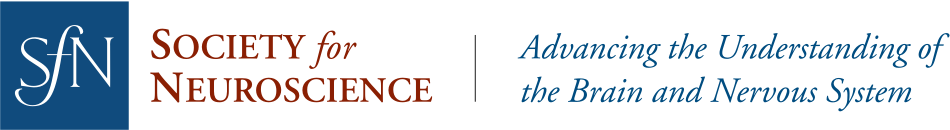Novel Viral-Genetic Method for Tracing Axon Collaterals of Broadly Projecting Neurons
 |
|
|
Material below summarizes the article An Intersectional Viral-Genetic Method for Fluorescent Tracing of Axon Collaterals Reveals Details of Noradrenergic Locus Coeruleus Structure, published on April 30, 2020, in eNeuro and authored by Nicholas W. Plummer, Daniel J. Chandler, Jeanne M. Powell, Erica L. Scappini, Barry D. Waterhouse and Patricia Jensen.
Highlights
- TrAC (Tracing Axon Collaterals) is a new viral-genetic method that allows simultaneous visualization of axon collaterals from a genetically defined neuronal population and a projection-based subpopulation.
- TrAC was used to show that norepinephrine (NE)-containing locus coeruleus (LC) neurons projecting to the medial prefrontal cortex and primary motor cortex differ from each other and from the LC as a whole in their pattern of axon collateralization.
- Labeled LC-NE neurons have dense axon projections to their primary target in the cerebral cortex, and widespread, albeit sparse, collateral projections to other cortical and subcortical regions.
Study Question
How can we use viral-genetic strategies to analyze axon collateral distribution of neuronal subpopulations? Does the broadly projecting LC-NE system contain neuronal subpopulations that differ in their axonal projection patterns?
How This Research Advances What We Know
Understanding how neurons in the mammalian brain coordinate complex behaviors requires comprehensive knowledge of the brain regions that receive their axonal projections. The noradrenergic locus coeruleus (LC), for instance, projects to much of the central nervous system through a vast network of axons. However, investigations into the organizational principles of this efferent projection system have yielded conflicting results. While some studies have provided evidence for a modular LC with distinct subpopulations of neurons that each project to a small number of brain regions, others have reported highly branched LC axons that project to many functionally unrelated targets. To further investigate this problem, we developed TrAC, a method that allows us to visualize LC neurons projecting to a specific target site, including all of their collateral projections to other targets within the central nervous system.
Experimental Design or Methodology
To apply TrAC to the LC-NE system, we bred genetically modified mice in which LC-NE neurons are labeled with a red fluorescent protein (tdTomato), but capable of switching to green fluorescent protein (EGFP) in the presence of Cre recombinase. We injected the mice with a retrogradely transported canine adenovirus construct encoding Cre in either the medial prefrontal cortex (mPFC) or primary motor cortex (M1), cortical regions with distinct roles in regulation of executive functions and motor behaviors, respectively. Consequently, LC-NE neurons projecting to the injection site were labeled with EGFP, including all of their axon collaterals, while all other LC-NE neurons were labeled with tdTomato. We counted EGFP- and tdTomato-labeled cell bodies in the LC (n=4 mice/injection site) and calculated the density of labeled axons in a variety of cortical and subcortical regions (n=4 mice mPFC injection, n=3 mice M1 injection).
Results
We found that LC-NE neurons projecting to the mPFC were distributed throughout the rostrocaudal extent of the LC but constituted a greater percentage of the rostral LC than the caudal. LC-NE neurons projecting to M1 were likewise observed throughout the LC but were more evenly distributed along the rostral-caudal axis. Axons of both mPFC- and M1-projecting LC-NE neurons were widely distributed in the cortex and basal forebrain, with the highest percentage of labeled axons close to the virus injection site and sparser collateral projections at other sites. Compared with the mPFC- and M1-projecting subpopulations, we observed that the LC projects more uniformly to cortical and subcortical regions.
Interpretation
This study confirms the utility of TrAC for mapping axon collaterals of genetically defined neuronal subtypes. Unlike previous viral-genetic techniques, only a single viral injection is required for differentially labeling a projection-based subpopulation, rendering TrAC applicable to anatomically scattered, as well as compact neuronal populations. Our analysis of LC-NE neurons shows that mPFC- and M1-projecting subpopulations differ from each other and from the full LC in their patterns of axon collateralization. These results are consistent with prior studies indicating that LC-NE neurons tend to have a major target field, which they innervate densely, and widespread minor targets containing fewer axon collaterals. Future studies will examine the functional relationships between major and minor targets of LC axons (e.g. cognitive, motor, and sensory circuitries). As such the focus will be on the ability of LC output to coordinately regulate activity within and across neural networks to achieve specific behavioral outcomes.
Visit eNeuro to read the original article and explore other content. Read other summaries of eNeuro and JNeurosci papers in the Neuronline collection SfN Journals: Research Article Summaries.
An Intersectional Viral-Genetic Method for Fluorescent Tracing of Axon Collaterals Reveals Details of Noradrenergic Locus Coeruleus Structure. Nicholas W. Plummer, Daniel J. Chandler, Jeanne M. Powell, Erica L. Scappini, Barry D. Waterhouse and Patricia Jensen. eNeuro 30 April 2020, 7 (3) 0010-20.2020; DOI: 10.1523/ENEURO.0010-20.2020