Distinct Genetic Signatures of Human Cortical and Subcortical Memory
Material below summarizes the article Distinct Genetic Signatures of Cortical and Subcortical Regions Associated with Human Memory, published on December 9, 2019, in eNeuro and authored by Pin Kwang Tan, Egor Ananyev, and Po-Jang Hsieh.
Highlights
- Despite a common involvement in memory, gene signatures of cortical and subcortical memory are largely distinct.
- Cortical memory genes are linked to immune and epigenetic regulation, while subcortical memory genes are associated with neurogenesis and the differentiation of glial cells.
- Overlapping genes between cortical and subcortical memory are involved in transcription regulation, synaptic plasticity, and glutamate receptor signaling.
 |
|
|
Study Question
We asked if there are genes associated with cortical and subcortical memory regions in the adult human brain. If so, as cortical and subcortical regions share a common involvement in memory, do they have a similar gene signature?
How This Research Advances What We Know
Memory function is crucial for everyday life, ranging from spatial navigation to information retention. The anatomical and functional aspects of human memory are well characterized, owing to neuroimaging and case studies of brain damage. However, despite the knowledge that memory ability is heritable, a comprehensive view of the genetics of human memory remains lacking.
Here, we analyzed the three-dimensional spatial distribution of gene expression across memory regions to identify such gene signatures of human memory. Despite a common involvement in memory, we found that cortical and subcortical areas have largely distinct genetic signatures. Genes shared between cortical and subcortical signatures are involved in regulating transcription and synaptic plasticity, hinting at fundamental processes in memory. We further identified genes with novel links to memory, such as neurogenesis-related MIS18BP1. Overall, our work provides an unprecedented insight into biological processes and genes associated with memory, and proposes candidates for further experimental investigation.
Experimental Design or Methodology
To identify genes associated with general memory, we first applied a spatial correlation method on a gene expression map (Allen Human Brain Atlas) and a neuroimaging map (Neurosynth memory map), separately for cortical and subcortical memory areas. For each gene, this correlates gene expression level and z-scores of the memory map across the brain, where a high z-score suggests that the brain area is highly relevant for memory (e.g., hippocampus). Genes with a high correlation value are likely related to memory. This produced a gene list of ~16k genes and their correlation values per analysis.
To identify genes and biological processes related to cortical and subcortical memory in an unsupervised manner, we used bioinformatics tools to identify candidates from the gene lists above. To identify cortex- and subcortex-specific genes, we identified non-overlapping genes between their profiles. We validated our results by verifying that the memory gene signatures agreed with animal literature.
Results
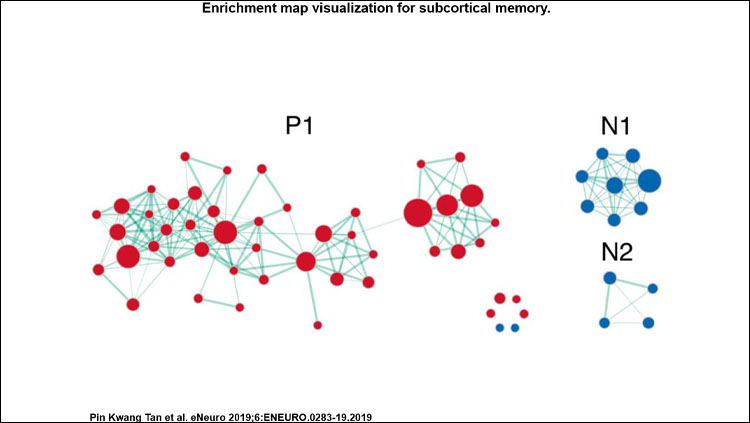
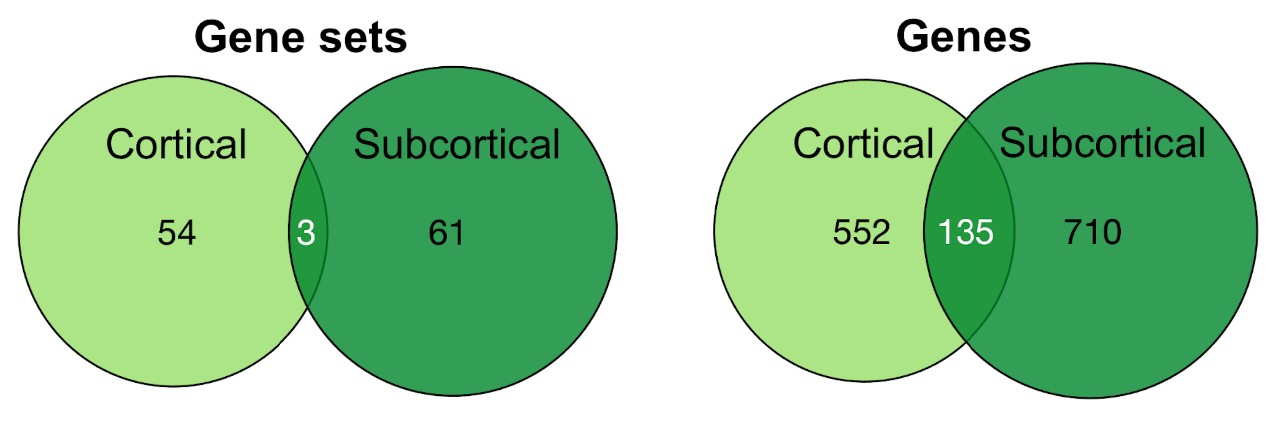
We found that the cortical and subcortical areas possess largely distinct gene signatures. There was no overlap in the top 10 cortical and subcortical memory genes, with a small degree overlap for all memory genes (9.6% out of 1,397 genes) and all biological process gene sets (2.5% out of 118 gene sets).
At the biological process level, we found differences in cortical and subcortical memory. Cortical processes include epigenetic regulation and immune signaling. Immune signaling received recent interest as an important factor in the onset and development of dementia. Subcortical processes include neurogenesis and glial cell differentiation. Dysfunctional glial-mediated potassium homeostasis and myelination was recently linked to memory deficits.
At the gene level, enriched genes for cortical and subcortical memory were similarly distinct. For instance, the top 10 candidate genes of PRKCD and CDK5 are linked to cortical and subcortical regions respectively. Interestingly, a small proportion of all memory genes (9.6%, or 135 genes) were shared between cortical and subcortical regions. These genes are linked to the Arp2/3 complex and the GABA and AMPA ligand-gated ion channels, which are relevant for memory function. Future work may look into how the convergence and divergence between their genetic profiles enable cortico-subcortical-specific functions in memory.
We found that the method was highly effective, as the probability of obtaining the number of memory genes observed was significantly above chance. The method is also specific, as it identifies more genes associated with the cognitive function analysed than for the control function (e.g., memory and motor function).
Interpretation
We show for the first time that cortical and subcortical regions involved in human memory possess distinct genetic signatures. These gene signatures are in agreement with prior research and were dissociable from a control function (motor function). Lesser-understood biological processes and genes may guide future experimental investigations in human cortical and subcortical memory function and disorders.
Presently, human memory evidence is mainly derived from popular non-invasive methods such as GWAS, which identifies gene variants related to brain or behavioral measures of cognition. However, these methods do not account for the spatial pattern of gene expression in the brain. Our approach relies on this spatial pattern and identifies genetic signatures related to human memory. Crucially, our human-centric, unsupervised approach is versatile, as it can reveal insights into any cognitive functions that are clinically relevant but with a genetic basis that is less understood (e.g., language, as in the case of dyslexia).
Visit eNeuro to read the original article and explore other content. Read other summaries of eNeuro and JNeurosci papers in the Neuronline collection SfN Journals: Research Article Summaries.
Distinct Genetic Signatures of Cortical and Subcortical Regions Associated with Human Memory. Pin Kwang Tan, Egor Ananyev, and Po-Jang Hsieh. eNeuro 9 December 2019, 6 (6) 0283-19.2019; DOI: 10.1523/ENEURO.0283-19.2019.